The Repartition Join we saw in the previous part is a Reduce-Side Join, because the actual joining is done in the reducer. The Replicated Join we are going to discover in this post is a Map-Side Join. The joining is done in mappers, and no reducer is even needed for this operation. So in a sense it should be faster join, but only if certain requirements are met.
We will see how it works and try using it on our Donations/Projects datasets and see if we can accomplish the same join we did in the previous post.
Contents
The Replicated Join
Basic Idea and Example
The main idea in the Replicated Join is to cache all data from the smaller dataset into the mappers which process splits of the bigger dataset.
In our example, our Donations sequence file is 4-blocks long, but our Projects sequence file is only 1-block long. The aim is to cache the Projects data in each of the 4 Donations mappers. This way, each mapper has all the Project records in memory and can use them to join its Donation records from the input split.
Here is an illustration, using the same sample data as in this Repartition Join illustration from the previous part. Once again, for simplicity we only show 2 Donation mappers, each input split contains only 3 records, and IDs in the input split keys have been simplified to “donXX” for Donations and “proXX” for Projects.
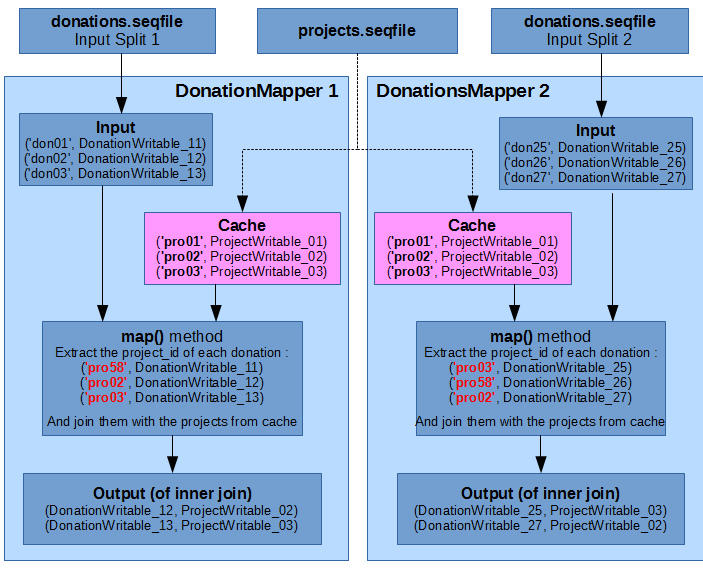
There is no need for reducers in this case, because the join results are already available in the outputs of the mappers.
However the results are not grouped in the same way as when using the Repartition Join from the previous post :
- When using a Repartition join (previous part), the join results are grouped by the same joining key in a given reducer output file.
- When using a Replicated join, the results are grouped by the bigger dataset records. The mapper outputs still contain the same records as their respective input splits.
For example, when looking at the illustration from my previous post on the Repartition Join, we can see that the join results involving the joining key “pro_02” are all in the same reducer output file at the end. The left side of these results are Donation records which come from 2 different input splits.
However in this Replicated Join illustration using the same datasets, we can see that the results involving the same joining key “pro_02” are separated across both mappers. But each mapper has in its output only the same Donation records which were in their original input split.
Requirements and Limitations
There are 2 important limitations using the Replicated Join:
- The smallest dataset has to fit into memory.
- Only an Inner Join or a Left Join can be performed.
The first point is pretty self-explanatory. Just keep in mind that our Projects dataset is only 1 HDFS block (< 128 MB), but it is compressed. The uncompressed sequence file is around 365 MB. And when loading this serialized data into memory, it will be a lot bigger in the JVM, because Java object memory footprints are huge, especially Strings objects which are at least 40 bytes each.
The second point needs a bit more discussion.
In our case we used the smaller Projects dataset as the Right side of the join. In each mapper what we have is all Project records, but only a subset of all Donations records. So given a Donation record (left side record), we can be sure that it can be joined to all Project records (right side records) matched by joining key, within the same mapper. For each Donation record (left side), if a mapper finds no matching Project record (right side), it can be sure that there is no matching record in any of the other mappers, since they all have the same cached data. So when finding no right side record, it can choose not to output anything (which produces an Inner Join), or it can choose to still output the left side record with a “null” right side (which produces a Left Join).
On the other hand, if a mapper finds no matching Donation record (left side) for a given Project record (right side), then it doesn’t know if any other mapper has a match, because each mapper has a different split of Donation records. This makes it impossible to make a Right Join. Let’s say it still wants to perform a Right Join, and believes that there is no matching left side, and outputs (null, Project_123). But maybe another mapper, when going through the same donation record, will find a matching left side Donation record, and output (Donation_456, Project_123). When merging both mapper outputs, we will find 2 inconsistent results. And in the case of all mappers finding no matching left side, there will be result duplicates. Each mapper will print out the same (null, Project_123) in their output. Because of this it is not possible to perform a consistent Full Outer Join either using a Replicated Join.
Basic Implementation
Here is the code for a basic Replicated Join : https://github.com/nicomak/[…]/ReplicatedJoinBasic.java
The important things to do are :
- Adding the projects dataset to the distributed cache in the driver code.
- Overriding the mapper’s setup(Context context) method to set up a cache map before the real map process begins.
- In the map method, for each Donation record, check the cache map to find any matching Project record.
- Explicitly setting the number of reduce tasks to 0 to suppress the whole reduce phase (otherwise the default Reducer will be used).
Job Execution and Analysis
Let’s run this job and see what happens :
$ hadoop jar donors.jar mapreduce.join.replicated.ReplicatedJoinBasic donors/donations.seqfile donors/projects.seqfile donors/output_replicatedjoinbasic
16/02/03 17:04:26 INFO client.RMProxy: Connecting to ResourceManager at ubuntu0/192.168.0.200:8032
16/02/03 17:04:29 INFO input.FileInputFormat: Total input paths to process : 1
16/02/03 17:04:30 INFO mapreduce.JobSubmitter: number of splits:4
16/02/03 17:04:32 INFO mapreduce.Job: The url to track the job: http://ubuntu0:8088/proxy/application_144894396118_0184/
16/02/03 17:04:32 INFO mapreduce.Job: Running job: job_1448943961184_0184
16/02/03 17:04:53 INFO mapreduce.Job: Job job_1448943961184_0184 running in uber mode : false
16/02/03 17:04:53 INFO mapreduce.Job: map 0% reduce 0%
16/02/03 17:08:36 INFO mapreduce.Job: Task Id : attempt_1448943961184_0184_m_000002_0, Status : FAILED
Container killed on request. Exit code is 137
Container exited with a non-zero exit code 137
Killed by external signal
16/02/03 17:08:37 INFO mapreduce.Job: map 0% reduce 0%
16/02/03 17:10:24 INFO mapreduce.Job: Task Id : attempt_1448943961184_0184_m_000002_1, Status : FAILED
Container [pid=27340,containerID=container_1448943961184_0184_01_000006] is running beyond physical memory limits.
Current usage: 1.0 GB of 1 GB physical memory used; 1.3 GB of 2.1 GB virtual memory used. Killing container.
Dump of the process-tree for container_1448943961184_0184_01_000006 :
|- PID PPID PGRPID SESSID CMD_NAME USER_MODE_TIME(MILLIS) SYSTEM_TIME(MILLIS) VMEM_USAGE(BYTES) RSSMEM_USAGE(PAGES) FULL_CMD_LINE
Oops ! The job has failed ! And we can see in the output that our 1 GB of allocated memory have been fully used. As I explained earlier, this is because the deserialized Writable objects are too big to all fit into memory.
Let’s try and estimate how much memory we need to cache all the ProjectWritable objects :
- The dataset contains approx. 880k records.
- Consider the following basic object sizes in Java 64-bit:
- String : [36 + length*2] rounded to a multiple of 8 ⇒ Let’s say approx. 56 bytes for 10-char strings.
- float primitive : 4 bytes
- Each ProjectWritable object contains :
- 15 String attribute objects : 15*56 = 840 bytes
- 3 float attributes : 3*4 = 12 bytes
- 15 references to the string attributes : 15*8 = 120 bytes
- So the total memory footprint for each record is around 972 bytes per object.
- So we need 880,000*972 = 855 MiB = approx. 816 MB for all records .
- Kept inside a map, which has a string object for each key, the total size of the map is then :
- The ProjectWritable objects : 816 MB
- The references to those writables : 880,000 * 8 = 7,04 MiB = approx. 6.7 MB
- The uid (32-char long) String objects used as keys : 880,000 * 104 = 91.5 MiB = approx. 87 MB
- The references to those String map keys : 880,000 * 8 = 7,04 MiB = approx. 6.7 MB
- So the total memory used by our map cache would be : 916 MB.
Our container is allocated around 1 GB of memory, of which a bit less than 900 MB is reserved for the JVM. And we’re out of luck because our map cache alone needs more than 900 MB of memory. That is why the job failed.
We have 2 choices : increase the memory allocation of our container, or optimize how we manage our memory. The first option is not interesting, and not always possible, so let’s get started with the optimizations !
Optimizations
Data Projection
We can project the data and make it smaller by selecting only the necessary fields before putting them into the cache. We will use the same two methods that we used in the previous post with the Repartition Join :
- Using smaller objects : this time they don’t need to be Writable hadoop objects. They can be simple POJOs because they won’t we serialized for shuffling since there is no reduce phase.
- Using manually “serialized” strings containing pipe-separated values. We can use normal String objects instead of hadoop Text as well because there is no shuffling.
Using smaller Objects
Here is the code for the same job, but which minimized space by using smaller objects in the cache map: https://github.com/nicomak/[…]/ReplicatedJoinProjectedByObjects.java
The job is successful this time :
$ hadoop jar donors.jar mapreduce.join.replicated.ReplicatedJoinProjectedByObjects donors/donations.seqfile donors/projects.seqfile donors/output_replicatedjoinprojectedobjects hdfs dfs -ls -h donors/output_repartitionjoinprojectedwritables Found 5 items -rw-r--r-- 2 hduser supergroup 0 2016-02-03 18:47 donors/output_replicatedjoinprojectedobjects/_SUCCESS -rw-r--r-- 2 hduser supergroup 196.9 M 2016-02-03 18:44 donors/output_replicatedjoinprojectedobjects/part-m-00000 -rw-r--r-- 2 hduser supergroup 197.0 M 2016-02-03 18:46 donors/output_replicatedjoinprojectedobjects/part-m-00001 -rw-r--r-- 2 hduser supergroup 196.3 M 2016-02-03 18:47 donors/output_replicatedjoinprojectedobjects/part-m-00002 -rw-r--r-- 2 hduser supergroup 167.7 M 2016-02-03 18:44 donors/output_replicatedjoinprojectedobjects/part-m-00003 $ hdfs dfs -cat donors/output_replicatedjoinprojectedobjects/part-m-00001 | head 435da4114b93084ffde6f459a68c86a0|341d7826c1fcfcf6712d1a9d5a1503f5||2010-01-21 20:57:38.946|10.00 341d7826c1fcfcf6712d1a9d5a1503f5|Washington|highest poverty|Health & Life Science 435de4e0d5df220b0a3eb02c6d387afb|5d0833a4363dafc1219fd792207ba194||2011-12-16 19:50:05.148|100.00 5d0833a4363dafc1219fd792207ba194|Dublin|moderate poverty|Environmental Science
The total size of the 4 mapper outputs is 196.9 + 197.0 + 196.3 + 167.7 = 757.9 MB.
Using the optimized Repartition join of the previous post, the total size of the 3 reducer outputs when performing the same Inner Join was 252.1+254.4+251.4 = 757.9 MB.
This confirms that we have exactly the same results using the Replicated Join and the Repartition Join.
The execution time was of 5 min 23 sec on average.
Using Strings
Instead of waiting for the map() method to print out the pipe-separated text result, we can directly store the text result in the cache in the setup method.
Here is the code for this job : https://github.com/nicomak/[…]/ReplicatedJoinProjectedByStrings.java
This job took only 3 min 30 sec on average.
Further optimization using a Semi-Join
Until now, what we did is to put all records from the small dataset in a cache. And to save space in the cache, we discarded unused attributes. But what if we could know which Project records are completely useless (i.e. will never be joined to a record from the big dataset) before creating the cache ? That could save a lot of space by discarding entire useless rows of data.
Let’s consider that we are performing an Inner Join between both datasets, with conditions on both sides (data must be filtered on both sides before the join).
A sort of Semi-Join can be performed in 3 steps to optimize memory usage:
- Create a simple MapReduce job to process the biggest dataset, donations.seqfile, applying all its WHERE clause conditions, and output all distinct values of its foreign key field “project_id”. The output of this operation is a list of all project ids which will effectively be used for joining. Let’s call this output filtered_project_ids.txt.
- Perform the following replicated join:
- Left side (input splits): projects.seqfile
- Right side (cache): filtered_project_ids.txt
- Output : useful_projects.seqfile which is a subset of projects.seqfile but containing only project records which:
- Satisfy the conditions on “projects” (we can filter the left-side dataset during this job).
- Satisfy the conditions on “donations” for donation records to which they would be joined (pre-filtered in the right side thanks to Step 1).
- Do the real Replicated Join job using the newly filtered projects data file:
- Left side (input splits): donations.seqfile
- Right side (cache): useful_projects.seqfile
The first 2 steps are equivalent to doing a Left Semi-Join in SQL, such as:
SELECT * FROM projects p LEFT SEMI JOIN donations d ON (d.project_id = p.project_id) WHERE d.total > 100 AND p.school_state = 'NY';
Which is also equivalent to these queries:
-- Using "EXISTS"
SELECT * FROM projects p
WHERE EXISTS(
SELECT * FROM donations d
WHERE d.project_id = p.project_id
AND d.amount > 100
AND p.school_state = 'NY'
);
-- Using "IN"
SELECT * FROM projects p
WHERE project_id IN (
SELECT d.project_id FROM donations d
WHERE d.amount > 100
AND p.school_state = 'NY'
);
The result of this two-step semi-join is a fully-filtered projects dataset. It enables us to know ahead of time which right-side records will be used for an Inner Join, based on the query conditions on both sides, before the real join happens.
This minimizes the projects data to be cached in the final step which performs the real join and maximizes its chance of fitting into memory.
Great information.
Thanks for sharing.